Peer-reviewed publications (Citations = 4261, H-index = 18; updated 26/4/5)
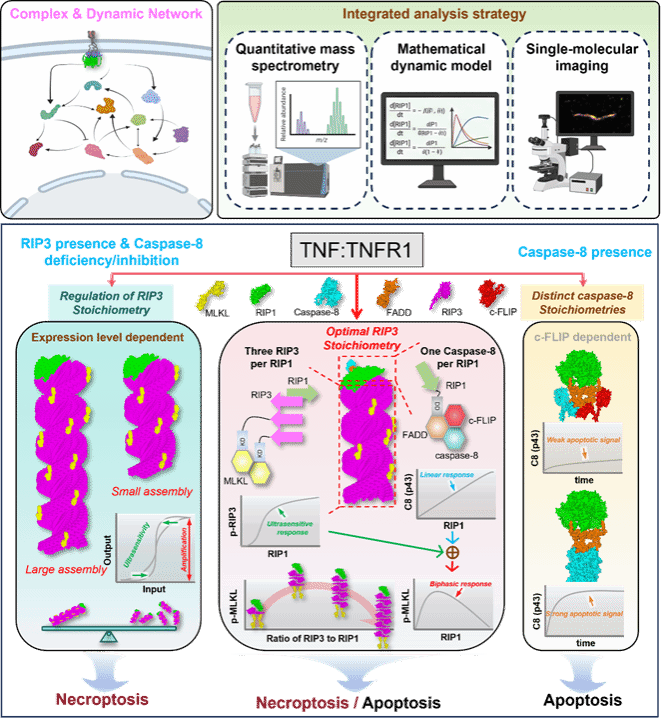
1. Li, X., Cao, Y., Xu, F., Zhang, Y., Kong, Y., Lan, C., Zhu, R., Lin, C., Zhong, C. Q., Liu, Z., Qi, H., Huang, Y., Xiao, Y., Sun, G. Q., Shuai, J., & Chen, X. (2025). Decoding necrosome assembly: harmonizing signal amplification and attenuation through optimal RIP3 stoichiometry. Nature communications, 17(1), 405. https://doi.org/10.1038/s41467-025-67098-5
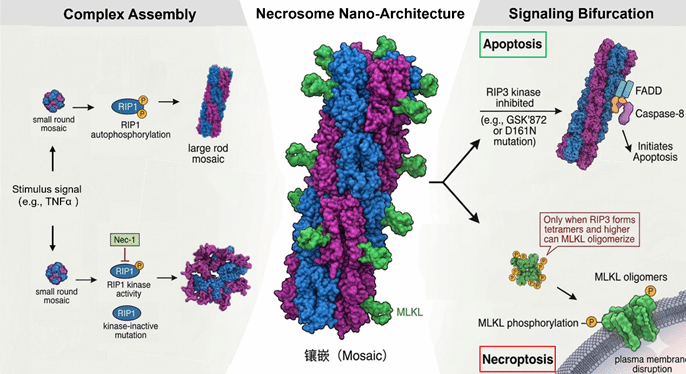
2. Chen, X., Zhu, R., Zhong, J., Ying, Y., Wang, W., Cao, Y., Cai, H., Li, X., Shuai, J., & Han, J. (2022). Mosaic composition of RIP1-RIP3 signalling hub and its role in regulating cell death. Nature cell biology, 24(4), 471–482. https://doi.org/10.1038/s41556-022-00854-7
3.Chen, X., He, W. T., Hu, L., Li, J., Fang, Y., Wang, X., Xu, X., Wang, Z., Huang, K., & Han, J. (2016). Pyroptosis is driven by non-selective gasdermin-D pore and its morphology is different from MLKL channel-mediated necroptosis. Cell research, 26(9), 1007–1020. https://doi.org/10.1038/cr.2016.100
4.Chen, X., Li, W., Ren, J., Huang, D., He, W. T., Song, Y., Yang, C., Li, W., Zheng, X., Chen, P., & Han, J. (2014). Translocation of mixed lineage kinase domain-like protein to plasma membrane leads to necrotic cell death. Cell research, 24(1), 105–121. https://doi.org/10.1038/cr.2013.171
5. Han, T., Ruan, C., Lin, H., Zhang, Y., Li, L., Sun, Y. H., Zhong, C. Q., Chen, X., Huang, K., Cao, Y., Fan, Z., Zhang, H., Han, J., & Zhang, Y. (2025). RIPK1 S161 phosphorylation promotes further autophosphorylation and cecal necroptosis in TNF-treated mice. The Journal of experimental medicine, 222(12), e20250277. https://doi.org/10.1084/jem.20250277
6. Li, X., Zhong, C. Q., Wu, R., Xu, X., Yang, Z. H., Cai, S., Wu, X., Chen, X., Yin, Z., He, Q., Li, D., Xu, F., Yan, Y., Qi, H., Xie, C., Shuai, J., & Han, J. (2021). RIP1-dependent linear and nonlinear recruitments of caspase-8 and RIP3 respectively to necrosome specify distinct cell death outcomes. Protein & cell, 12(11), 858–876. https://doi.org/10.1007/s13238-020-00810-x
7. Yang, Z., Wang, Y., Zhang, Y., He, X., Zhong, C. Q., Ni, H., Chen, X., Liang, Y., Wu, J., Zhao, S., Zhou, D., & Han, J. (2018). RIP3 targets pyruvate dehydrogenase complex to increase aerobic respiration in TNF-induced necroptosis. Nature cell biology, 20(2), 186–197. https://doi.org/10.1038/s41556-017-0022-y
8. Zhang, Y., Chen, X., Gueydan, C., & Han, J. (2018). Plasma membrane changes during programmed cell deaths. Cell research, 28(1), 9–21. https://doi.org/10.1038/cr.2017.133
9. Xue, Z., Wang, S., Li, J., Chen, X., Han, J., & Han, S. (2018). Bifunctional Super-resolution Imaging Probe with Acidity-Independent Lysosome-Retention Mechanism. Analytical chemistry, 90(19), 11393–11400. https://doi.org/10.1021/acs.analchem.8b02365
10. Zhang, Y., Su, S. S., Zhao, S., Yang, Z., Zhong, C. Q., Chen, X., Cai, Q., Yang, Z. H., Huang, D., Wu, R., & Han, J. (2017). RIP1 autophosphorylation is promoted by mitochondrial ROS and is essential for RIP3 recruitment into necrosome. Nature communications, 8, 14329. https://doi.org/10.1038/ncomms14329
11. Huang, D., Zheng, X., Wang, Z. A., Chen, X., He, W. T., Zhang, Y., Xu, J. G., Zhao, H., Shi, W., Wang, X., Zhu, Y., & Han, J. (2017). The MLKL Channel in Necroptosis Is an Octamer Formed by Tetramers in a Dyadic Process. Molecular and cellular biology, 37(5), e00497-16. https://doi.org/10.1128/MCB.00497-16
12. Wu, X. N., Yang, Z. H., Wang, X. K., Zhang, Y., Wan, H., Song, Y., Chen, X., Shao, J., & Han, J. (2014). Distinct roles of RIP1-RIP3 hetero- and RIP3-RIP3 homo-interaction in mediating necroptosis. Cell death and differentiation, 21(11), 1709–1720. https://doi.org/10.1038/cdd.2014.77
13. Zhuang, R., Zhou, Y., Wang, Z., Cao, Y., Chen, J., Xu, L., Ren, Y., Zheng, Y., Wei, Z., Qiu, H., Li, L., Han, Y., Yun, Y., Chen, X., Hong, W., & Wang, T. (2023). Rab26 restricts insulin secretion via sequestering Synaptotagmin-1. PLoS biology, 21(6), e3002142. https://doi.org/10.1371/journal.pbio.3002142
14. Peng, G. X., Mao, X. L., Cao, Y., Yao, S. Y., Li, Q. R., Chen, X., Wang, E. D., & Zhou, X. L. (2022). RNA granule-clustered mitochondrial aminoacyl-tRNA synthetases form multiple complexes with the potential to fine-tune tRNA aminoacylation. Nucleic acids research, 50(22), 12951–12968. https://doi.org/10.1093/nar/gkac1141
15. Jiang, B., Zhang, J., Zhao, G., Liu, M., Hu, J., Lin, F., Wang, J., Zhao, W., Ma, H., Zhang, C., Wu, C., Yao, L., Liu, Q., Chen, X., Cao, Y., Zheng, Y., Zhang, C., Han, A., Lin, D., & Li, Q. (2022). Filamentous GLS1 promotes ROS-induced apoptosis upon glutamine deprivation via insufficient asparagine synthesis. Molecular cell, 82(10), 1821–1835.e6. https://doi.org/10.1016/j.molcel.2022.03.016
16. Zhou, X., Ma, F., Xie, J., Yuan, M., Li, Y., Shaabani, N., Zhao, F., Huang, D., Wu, N. C., Lee, C. D., Liu, H., Li, J., Chen, Z., Hong, Y., Liu, W. H., Xiao, N., Burton, D. R., Tu, H., Li, H., Chen, X., … Huang, Z. (2021). Diverse immunoglobulin gene usage and convergent epitope targeting in neutralizing antibody responses to SARS-CoV-2. Cell reports, 35(6), 109109. https://doi.org/10.1016/j.celrep.2021.109109
17. Li, M., Zhang, C. S., Zong, Y., Feng, J. W., Ma, T., Hu, M., Lin, Z., Li, X., Xie, C., Wu, Y., Jiang, D., Li, Y., Zhang, C., Tian, X., Wang, W., Yang, Y., Chen, J., Cui, J., Wu, Y. Q., Chen, X., … Lin, S. C. (2019). Transient Receptor Potential V Channels Are Essential for Glucose Sensing by Aldolase and AMPK. Cell metabolism, 30(3), 508–524.e12. https://doi.org/10.1016/j.cmet.2019.05.018
18. Wang, P., Geng, J., Gao, J., Zhao, H., Li, J., Shi, Y., Yang, B., Xiao, C., Linghu, Y., Sun, X., Chen, X., Hong, L., Qin, F., Li, X., Yu, J. S., You, H., Yuan, Z., Zhou, D., Johnson, R. L., & Chen, L. (2019). Macrophage achieves self-protection against oxidative stress-induced ageing through the Mst-Nrf2 axis. Nature communications, 10(1), 755. https://doi.org/10.1038/s41467-019-08680-6
19. Gao, H., Yang, Z., Wang, X., Qian, P., Hong, R., Chen, X., Su, X. Z., Cui, H., & Yuan, J. (2018). ISP1-Anchored Polarization of GCβ/CDC50A Complex Initiates Malaria Ookinete Gliding Motility. Current biology : CB, 28(17), 2763–2776.e6. https://doi.org/10.1016/j.cub.2018.06.069
20. Chen, X., Ma, J. J., Tan, M., Yao, P., Hu, Q. H., Eriani, G., & Wang, E. D. (2011). Modular pathways for editing non-cognate amino acids by human cytoplasmic leucyl-tRNA synthetase. Nucleic acids research, 39(1), 235–247. https://doi.org/10.1093/nar/gkq763
